Validation Data Gallery
Tested Applications
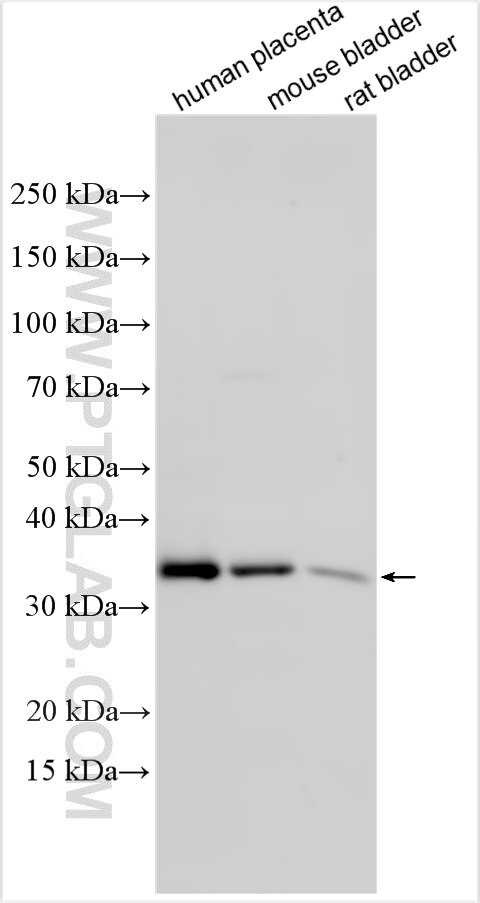
| Positive WB detected in | human placenta tissue, mouse bladder tissue, rat bladder tissue |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:2000-1:16000 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Product Information
55300-1-AP targets LIMS2 in WB, ELISA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
Peptide 相同性解析による交差性が予測される生物種 |
| Full Name | LIM and senescent cell antigen-like domains 2 |
| Calculated molecular weight | 39 kDa |
| Observed molecular weight | 35-40 kDa |
| GenBank accession number | NM_017980 |
| Gene Symbol | LIMS2 |
| Gene ID (NCBI) | 55679 |
| RRID | AB_3670247 |
| Conjugate | Unconjugated |
| Form | |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | Q7Z4I7 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol{{ptg:BufferTemp}}7.3 |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. |
Background Information
LIMS2, also known as PINCH2, is a member of a small family of focal adhesion proteins. LIMS2 contains five LIM domains, each domain forming two zinc fingers, which permit interactions that regulate cell shape and migration. Along with integrin-linked kinase and parvin, LIMS2 is a component of a complex that mediates multiple protein-protein interactions at adhesion sites between cells and the extracellular matrix (ECM) (PMID: 12167643). Mutations in LIMS2 are associated with a novel muscular dystrophy, severe cardiomyopathy and triangular tongues(PMID: 25589244).
Protocols
| Product Specific Protocols | |
|---|---|
| WB protocol for LIMS2 antibody 55300-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |