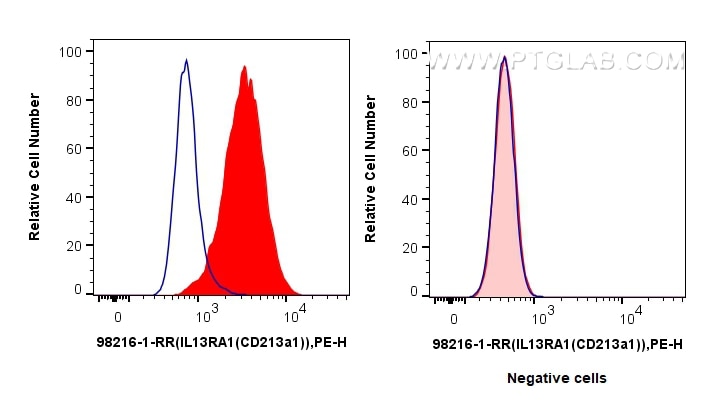
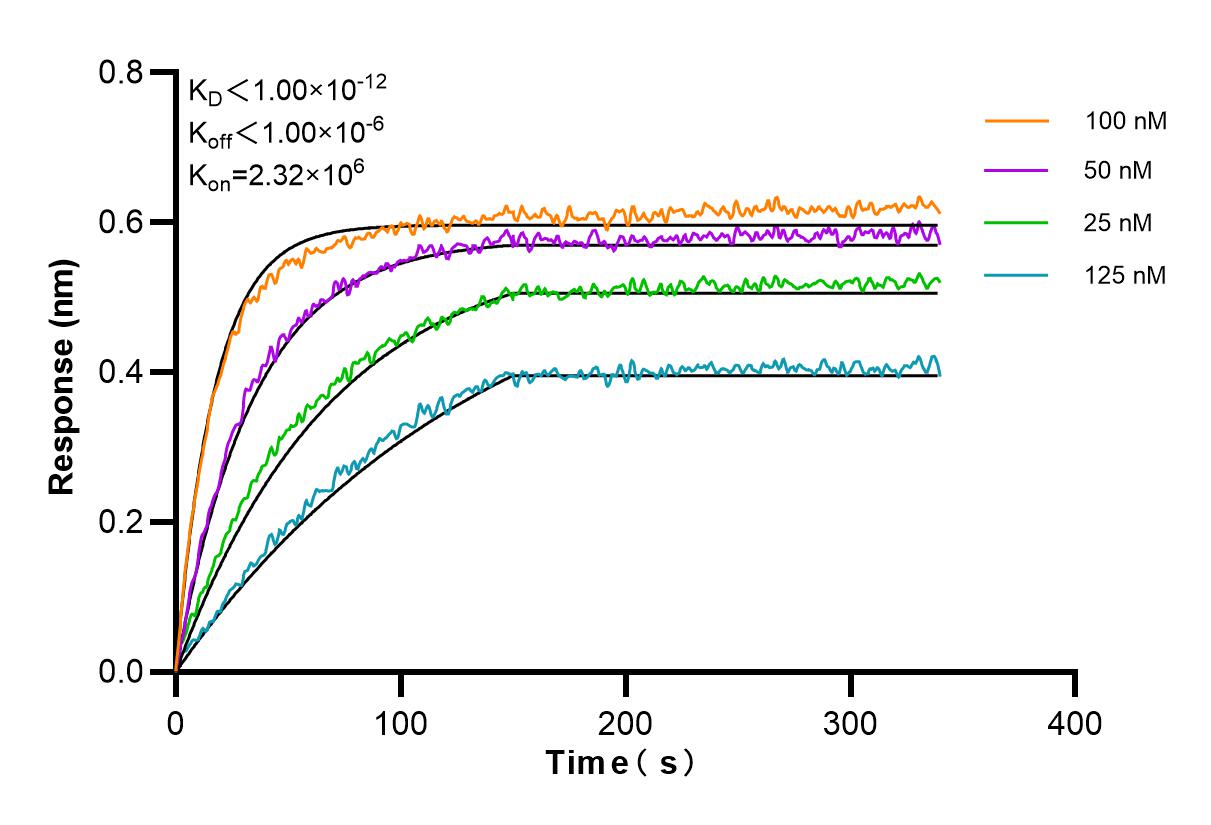
Validation Data Gallery
Tested Applications
| Positive FC detected in | MCF-7 cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Flow Cytometry (FC) | FC : 0.13 ug per 10^6 cells in 100 μl suspension |
| This reagent has been tested for flow cytometric analysis. It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Product Information
98216-1-RR targets IL-13Ra1/CD213a1 in FC applications and shows reactivity with human samples.
| Tested Reactivity | human |
| Host / Isotype | Rabbit / IgG |
| Class | Recombinant |
| Type | Antibody |
| Immunogen |
Recombinant protein 相同性解析による交差性が予測される生物種 |
| Full Name | interleukin 13 receptor, alpha 1 |
| Calculated molecular weight | 49 kDa |
| GenBank accession number | BC009960 |
| Gene Symbol | IL-13Rα1 |
| Gene ID (NCBI) | 3597 |
| Conjugate | Unconjugated |
| Form | |
| Form | Liquid |
| Purification Method | Protein A purfication |
| UNIPROT ID | P78552 |
| Storage Buffer | PBS with 0.09% sodium azide{{ptg:BufferTemp}}7.3 |
| Storage Conditions | Store at 2 - 8°C. Stable for one year after shipment. |
Background Information
IL-13 is an immunoregulatory cytokine secreted predominantly by activated T-helper 2 (Th2) cells and performs multiple functions through a complex receptor system comprised of IL-4Rα and two IL-13 binding proteins, IL-13Rα1 and IL-13Rα2 (PMID: 12704343). IL-13Rα1 is a glycosylated transmembrane protein (PMID: 10856136; 9723655). IL-13Rα1 is expressed on a wide range of cells including B cells, monocytes, macrophages, basophils, mast cells, and endothelial cells (PMID: 10856136; 31866859). IL-13 binding to the IL-4Rα/IL-13Rα1 complex leads to activation of JAK1, JAK2, and TYK2 and subsequent phosphorylation of STAT6 and STAT3 (PMID: 31866859).
Protocols
| Product Specific Protocols | |
|---|---|
| FC protocol for IL-13Ra1/CD213a1 antibody 98216-1-RR | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |