Validation Data Gallery
Tested Applications
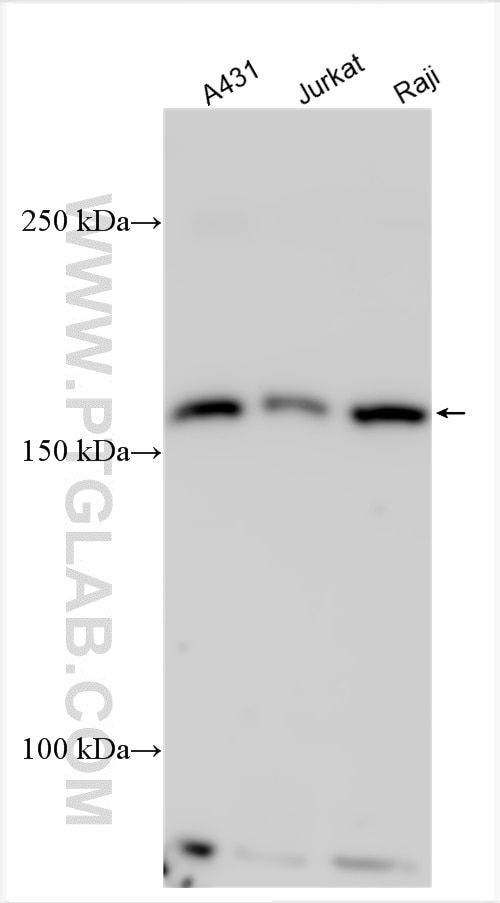
| Positive WB detected in | A431 cells, Jurkat cells, Raji cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:500-1:1000 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Product Information
22261-1-AP targets ESPL1 in WB, ELISA applications and shows reactivity with human samples.
| Tested Reactivity | human |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
Peptide 相同性解析による交差性が予測される生物種 |
| Full Name | extra spindle pole bodies homolog 1 (S. cerevisiae) |
| Calculated molecular weight | 233 kDa |
| Observed molecular weight | 170 kDa |
| GenBank accession number | NM_012291 |
| Gene Symbol | ESPL1 |
| Gene ID (NCBI) | 9700 |
| Conjugate | Unconjugated |
| Form | |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | Q14674 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol{{ptg:BufferTemp}}7.3 |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. |
Background Information
ESPL1 (Extra spindle poles-like 1 protein), also known as Separin, Separase, a cysteine endopeptidase, plays a vital role in the stable binding between sister chromatids before anaphase and their timely separation during anaphase, which is the key to chromosome inheritance (PMID: 34512713). ESPL1 expression is elevated and predicts poor outcomes in patients with various types of cancer, including breast cancer, glioma and endometrial cancer (PMID: 36465268). Full-length ESPL1 decreased after 30 min resulting in a ~ 170 kDa N-terminal (ESPL1N−term) and a ~ 60 kDa C-terminal polypeptide (ESPL1CC−term), reflecting ESPL1 activation and auto-cleavage (PMID: 30429481)
Protocols
| Product Specific Protocols | |
|---|---|
| WB protocol for ESPL1 antibody 22261-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |