Recombinant Human KLK14 protein (rFc Tag)(HPLC verified)
Species
Human
Purity
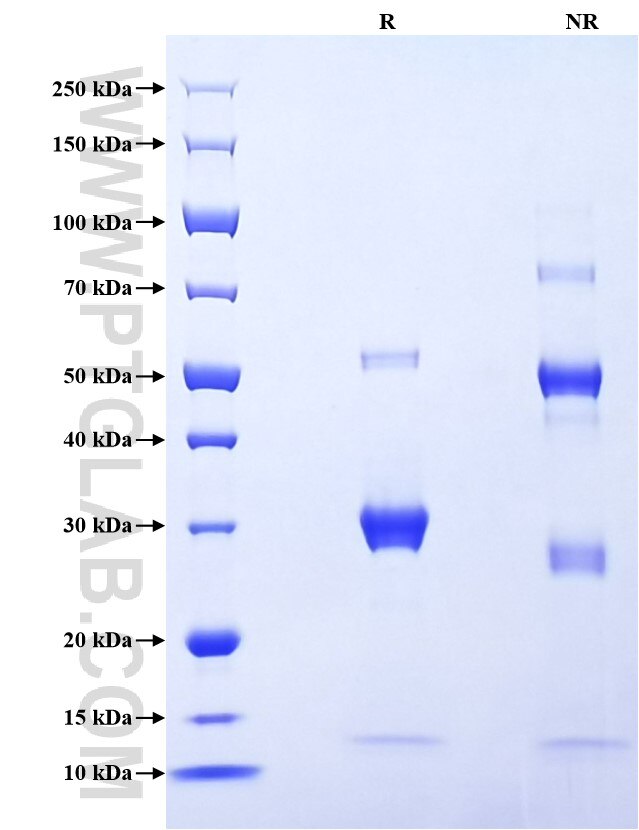
>90 %, SDS-PAGE
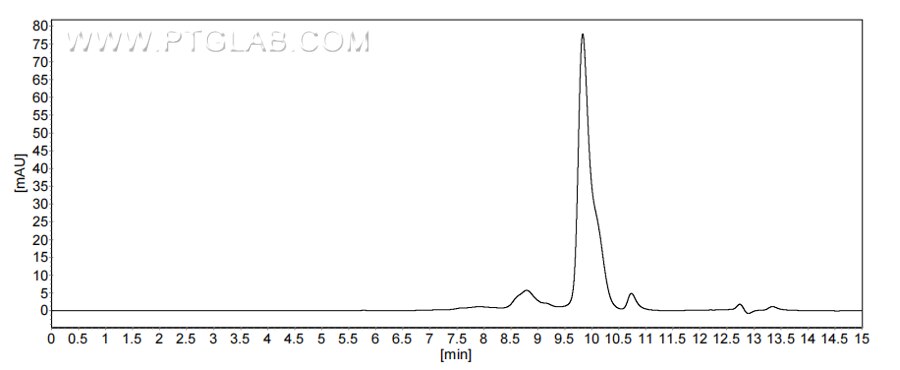
>90 %, SEC-HPLC
Tag
rFc Tag
Activity
not tested
Cat no : Eg2472
Validation Data Gallery
Product Information
| Purity | >90 %, SDS-PAGE >90 %, SEC-HPLC |
| Endotoxin | <0.1 EU/μg protein, LAL method |
| Activity |
Not tested |
| Expression | HEK293-derived Human KLK14 protein Gln35-Lys267 (Accession# Q9P0G3) with a rabbit IgG Fc tag at the C-terminus. |
| GeneID | 43847 |
| Accession | Q9P0G3 |
| PredictedSize | 51.5 kDa |
| SDS-PAGE | 13 kDa, 28-32 kDa and 55 kDa, reducing (R) condition |
| Formulation | Lyophilized from 0.22 μm filtered solution in PBS, pH 7.4. Normally 5% trehalose and 5% mannitol are added as protectants before lyophilization. |
| Reconstitution | Briefly centrifuge the tube before opening. Reconstitute at 0.1-0.5 mg/mL in sterile water. |
| Storage Conditions |
It is recommended that the protein be aliquoted for optimal storage. Avoid repeated freeze-thaw cycles.
|
| Shipping | The product is shipped at ambient temperature. Upon receipt, store it immediately at the recommended temperature. |
Background
KLK14, or Kallikrein-14, is a protein-coding gene that belongs to the kallikrein family of secreted serine proteases. Activation of KLK14 is mediated by KLK5, and once activated, KLK14 further amplifies the activity of KLK proteases through a positive feedback loop by cleaving pro-KLK5, which is a central player in the KLK cascade. One of the most notable substrates of KLK14 is PAR2 (Protease-Activated Receptor 2). KLK14 has been implicated in various physiological functions and is suggested to be involved in carcinogenesis, with potential as a novel biomarker for diseases such as cancer and skin disorders. It is associated with diseases like Netherton Syndrome and Ovarian Cancer. Moreover, KLK14 may play a role in the activation of membrane-type matrix metalloproteinases (MT-MMPs), which are involved in the proteolytic processing of components of the extracellular matrix, modulating the pericellular environment in physiology and pathologies.
References:
1. Katherine Falkowski . et al. (2020). Int J Mol Sci. 21(12):4383. 2. Hyesook Yoon. et al. (2007). J Biol Chem.282(44):31852-31864. 3. Hyun-Jeong Ra. et al. (2007). Matrix Biol. 26(8):587-596.