Validation Data Gallery
Tested Applications
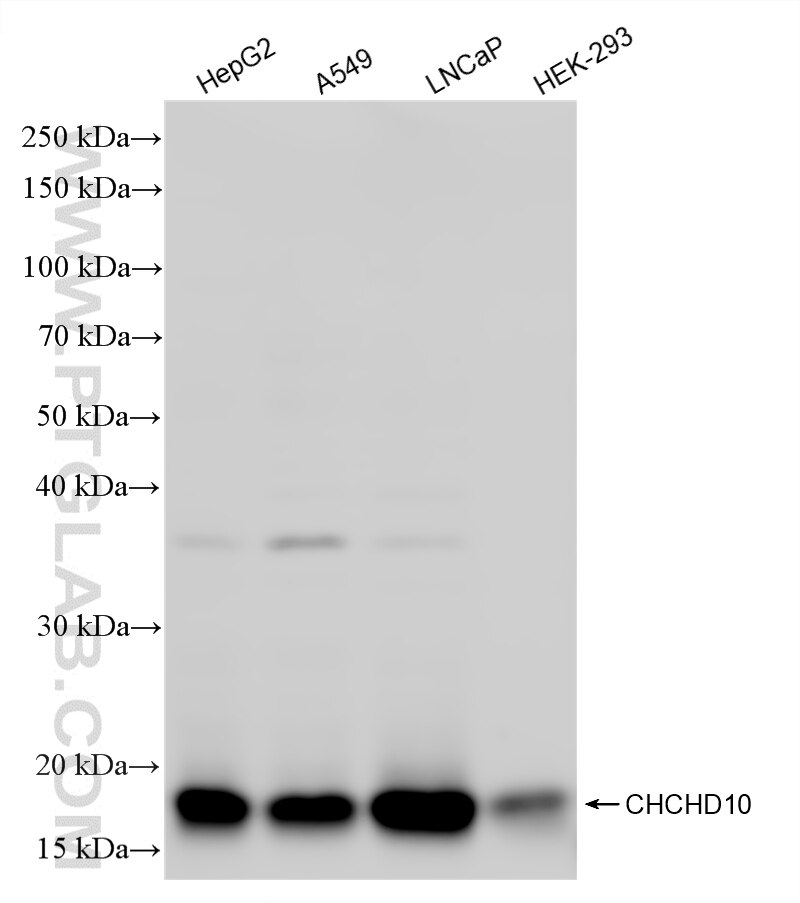
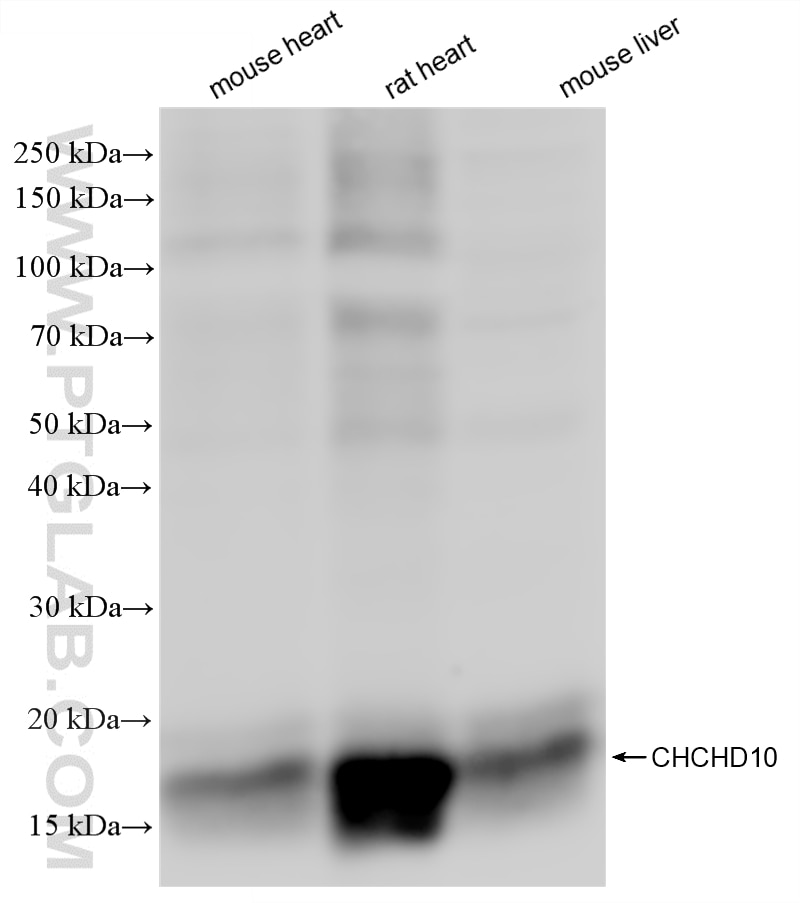
| Positive WB detected in | HepG2 cells, mouse heart tissue, A549 cells, LNCaP cells, HEK-293 cells, rat heart tissue, mouse liver tissue |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:500-1:2000 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Product Information
87812-3-RR targets CHCHD10 in WB, ELISA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Host / Isotype | Rabbit / IgG |
| Class | Recombinant |
| Type | Antibody |
| Immunogen |
CatNo: Ag22598 Product name: Recombinant human CHCHD10 protein Source: e coli.-derived, PET28a Tag: 6*His Domain: 82-142 aa of BC065232 Sequence: QPAVQQAPTPAAPQPLQMGPCAYEIRQFLDCSTTQSDLSLCEGFSEALKQCKYYHGLSSLP 相同性解析による交差性が予測される生物種 |
| Full Name | coiled-coil-helix-coiled-coil-helix domain containing 10 |
| Calculated molecular weight | 14 kDa |
| Observed molecular weight | 14-18 kDa |
| GenBank accession number | BC065232 |
| Gene Symbol | CHCHD10 |
| Gene ID (NCBI) | 400916 |
| Conjugate | Unconjugated |
| Form | |
| Form | Liquid |
| Purification Method | Protein A purification |
| UNIPROT ID | Q8WYQ3 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol{{ptg:BufferTemp}}7.3 |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. |
Background Information
CHCHD10 is a mitochondrial protein that is enriched at cristae junctions in the intermembrane space and is likely involved in mitochondrial genome stability and maintenance of cristae junctions. Mutations in CHCHD10 have recently been described as a cause of frontotemporal dementia (FTD) comorbid with amyotrophic lateral sclerosis (ALS).
Protocols
| Product Specific Protocols | |
|---|---|
| WB protocol for CHCHD10 antibody 87812-3-RR | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |